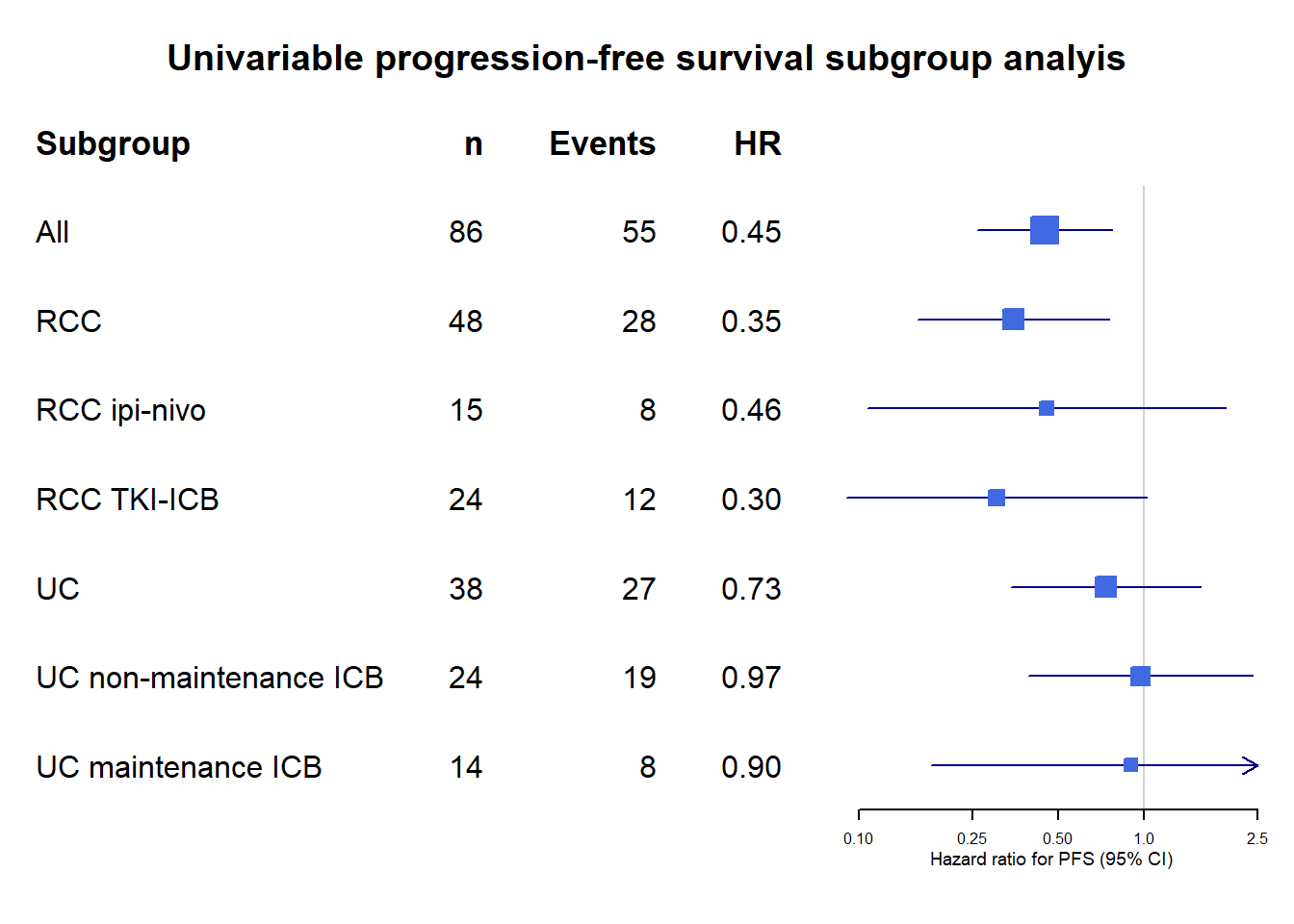
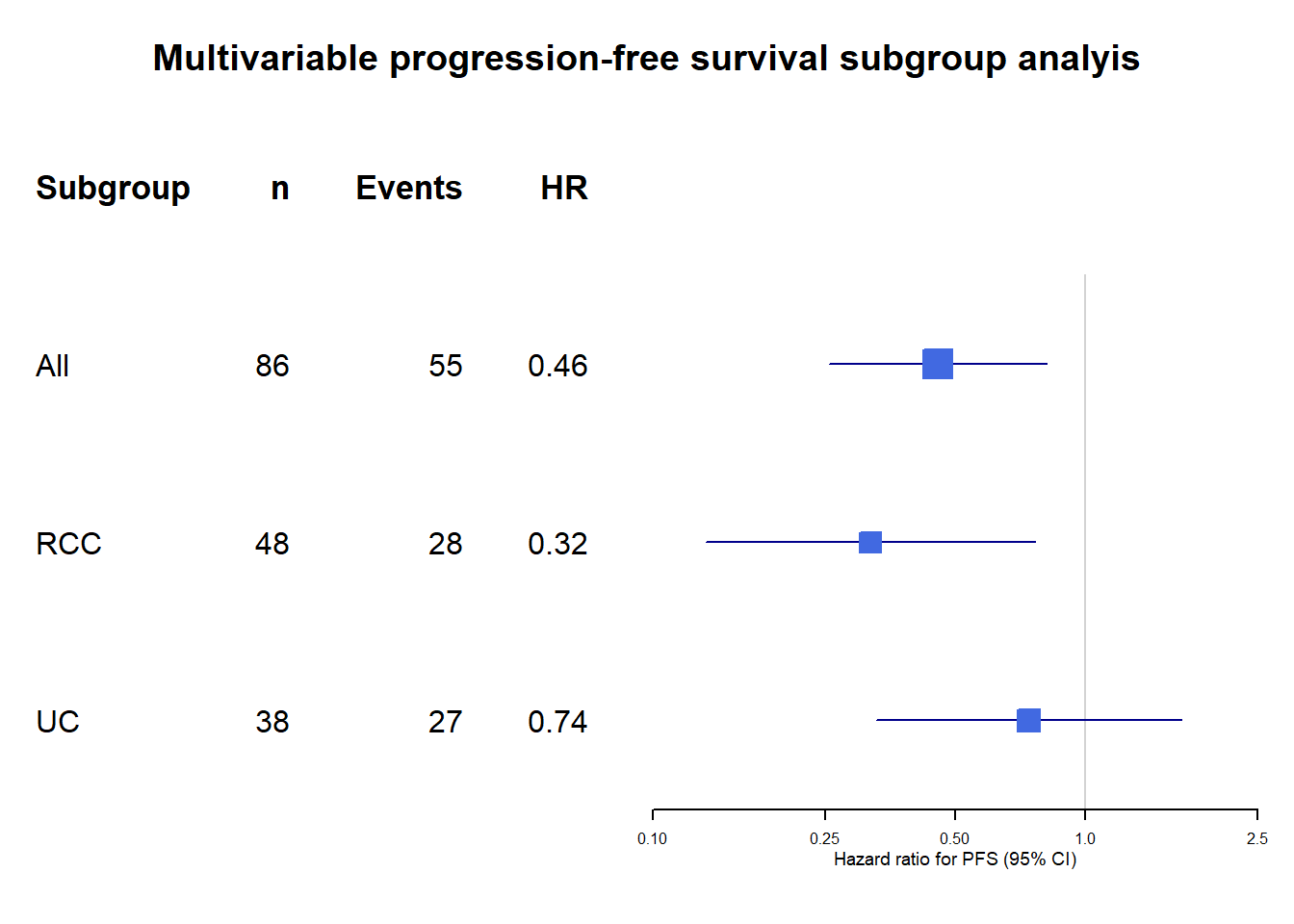
Univariate HRs for PFS among clinically relavant subgroups.
<- coxph (Surv (pfs_mo,pod_status)~ fiber_dichotomized,summary (cox_pfs_all_uv)
Call:
coxph(formula = Surv(pfs_mo, pod_status) ~ fiber_dichotomized,
data = all_metadata_fiber)
n= 86, number of events= 55
coef exp(coef) se(coef) z Pr(>|z|)
fiber_dichotomized -0.8011 0.4488 0.2780 -2.882 0.00395 **
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
exp(coef) exp(-coef) lower .95 upper .95
fiber_dichotomized 0.4488 2.228 0.2603 0.7739
Concordance= 0.606 (se = 0.033 )
Likelihood ratio test= 7.82 on 1 df, p=0.005
Wald test = 8.31 on 1 df, p=0.004
Score (logrank) test = 8.74 on 1 df, p=0.003
<- coxph (Surv (pfs_mo,pod_status)~ fiber_dichotomized,summary (cox_pfs_RC)
Call:
coxph(formula = Surv(pfs_mo, pod_status) ~ fiber_dichotomized,
data = mRCdataFiber)
n= 48, number of events= 28
coef exp(coef) se(coef) z Pr(>|z|)
fiber_dichotomized -1.0541 0.3485 0.3945 -2.672 0.00754 **
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
exp(coef) exp(-coef) lower .95 upper .95
fiber_dichotomized 0.3485 2.869 0.1608 0.7551
Concordance= 0.641 (se = 0.046 )
Likelihood ratio test= 6.46 on 1 df, p=0.01
Wald test = 7.14 on 1 df, p=0.008
Score (logrank) test = 7.8 on 1 df, p=0.005
<- coxph (Surv (pfs_mo,pod_status)~ fiber_dichotomized,summary (cox_pfs_RC_IpiNivo)
Call:
coxph(formula = Surv(pfs_mo, pod_status) ~ fiber_dichotomized,
data = dataRCIpiNivoFiber)
n= 15, number of events= 8
coef exp(coef) se(coef) z Pr(>|z|)
fiber_dichotomized -0.7864 0.4555 0.7387 -1.065 0.287
exp(coef) exp(-coef) lower .95 upper .95
fiber_dichotomized 0.4555 2.196 0.1071 1.937
Concordance= 0.627 (se = 0.094 )
Likelihood ratio test= 1.04 on 1 df, p=0.3
Wald test = 1.13 on 1 df, p=0.3
Score (logrank) test = 1.19 on 1 df, p=0.3
<- coxph (Surv (pfs_mo,pod_status)~ fiber_dichotomized,summary (cox_pfs_RC_TKI_IO)
Call:
coxph(formula = Surv(pfs_mo, pod_status) ~ fiber_dichotomized,
data = dataRCioTKIFiber)
n= 24, number of events= 12
coef exp(coef) se(coef) z Pr(>|z|)
fiber_dichotomized -1.1926 0.3034 0.6183 -1.929 0.0538 .
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
exp(coef) exp(-coef) lower .95 upper .95
fiber_dichotomized 0.3034 3.295 0.09031 1.02
Concordance= 0.635 (se = 0.073 )
Likelihood ratio test= 3.16 on 1 df, p=0.08
Wald test = 3.72 on 1 df, p=0.05
Score (logrank) test = 4.17 on 1 df, p=0.04
<- coxph (Surv (pfs_mo,pod_status)~ fiber_dichotomized,summary (cox_pfs_UC)
Call:
coxph(formula = Surv(pfs_mo, pod_status) ~ fiber_dichotomized,
data = mUCdataFiber)
n= 38, number of events= 27
coef exp(coef) se(coef) z Pr(>|z|)
fiber_dichotomized -0.3084 0.7346 0.3919 -0.787 0.431
exp(coef) exp(-coef) lower .95 upper .95
fiber_dichotomized 0.7346 1.361 0.3408 1.584
Concordance= 0.53 (se = 0.052 )
Likelihood ratio test= 0.61 on 1 df, p=0.4
Wald test = 0.62 on 1 df, p=0.4
Score (logrank) test = 0.62 on 1 df, p=0.4
<- coxph (Surv (pfs_mo,pod_status)~ fiber_dichotomized,summary (cox_pfs_UC_NOTavelumab)
Call:
coxph(formula = Surv(pfs_mo, pod_status) ~ fiber_dichotomized,
data = dataUC_NOTavelumabFiber)
n= 24, number of events= 19
coef exp(coef) se(coef) z Pr(>|z|)
fiber_dichotomized -0.03016 0.97029 0.46269 -0.065 0.948
exp(coef) exp(-coef) lower .95 upper .95
fiber_dichotomized 0.9703 1.031 0.3918 2.403
Concordance= 0.473 (se = 0.07 )
Likelihood ratio test= 0 on 1 df, p=0.9
Wald test = 0 on 1 df, p=0.9
Score (logrank) test = 0 on 1 df, p=0.9
<- coxph (Surv (pfs_mo,pod_status)~ fiber_dichotomized,summary (cox_pfs_UC_avelumab)
Call:
coxph(formula = Surv(pfs_mo, pod_status) ~ fiber_dichotomized,
data = dataUCavelumabFiber)
n= 14, number of events= 8
coef exp(coef) se(coef) z Pr(>|z|)
fiber_dichotomized -0.1060 0.8994 0.8239 -0.129 0.898
exp(coef) exp(-coef) lower .95 upper .95
fiber_dichotomized 0.8994 1.112 0.1789 4.521
Concordance= 0.527 (se = 0.091 )
Likelihood ratio test= 0.02 on 1 df, p=0.9
Wald test = 0.02 on 1 df, p=0.9
Score (logrank) test = 0.02 on 1 df, p=0.9
<- tibble:: tibble (mean = c (0.4488 ,0.3485 , 0.4555 , 0.3034 , 0.7346 , 0.97029 , 0.8994 ),lower = c (0.26032 ,0.1608 , 0.1071 , 0.09031 , 0.3408 , 0.3918 , 0.1789 ),upper = c ( 0.7739 ,0.7551 , 1.937 , 1.02 , 1.584 , 2.403 , 4.521 ),study = c ("All" ,"RCC" , "RCC ipi-nivo" , "RCC TKI-ICB" ,"UC" , "UC non-maintenance ICB" , "UC maintenance ICB" ),n = c ("86" ,"48" ,"15" ,"24" ,"38" ,"24" ,"14" ),events = c ("55" ,"28" ,"8" ,"12" ,"27" ,"19" ,"8" ),HR = c ("0.45" ,"0.35" , "0.46" , "0.30" , "0.73" , "0.97" , "0.90" ))|> forestplot (labeltext = c (study, n, events, HR),clip = c (0.09 , 2.5 ),xlog = TRUE ,xlab = "Hazard ratio for PFS (95% CI)" ,xticks = c (0.1 ,0.25 ,0.5 ,1.0 ,2.5 ),title = "Univariable progression-free survival subgroup analyis" ) |> fp_set_style (box = "royalblue" ,line = "darkblue" ,summary = "royalblue" ) |> fp_add_header (study = c ("Subgroup" ),n = c ("n" ),events = c ("Events" ),HR = c ("HR" ))
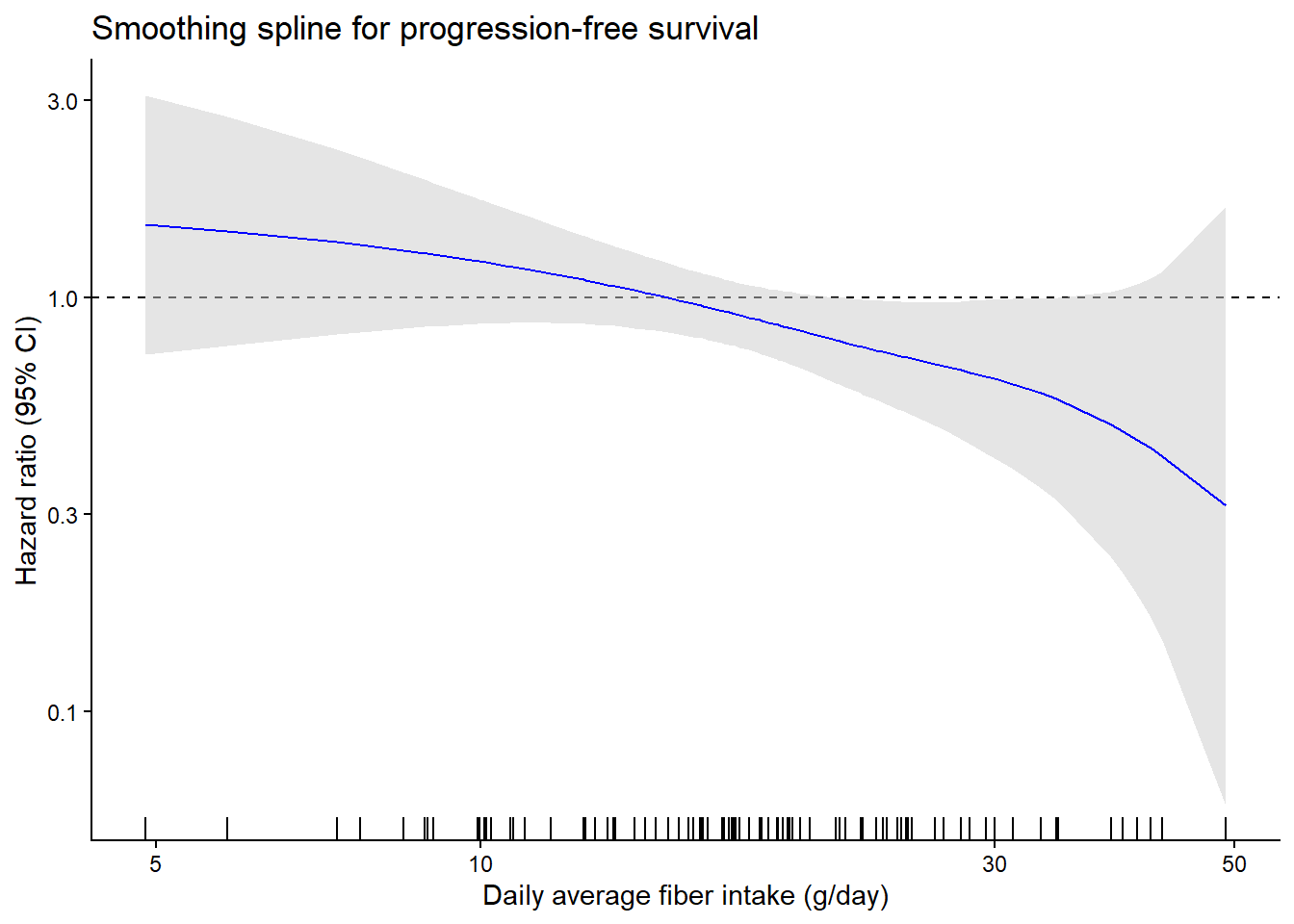
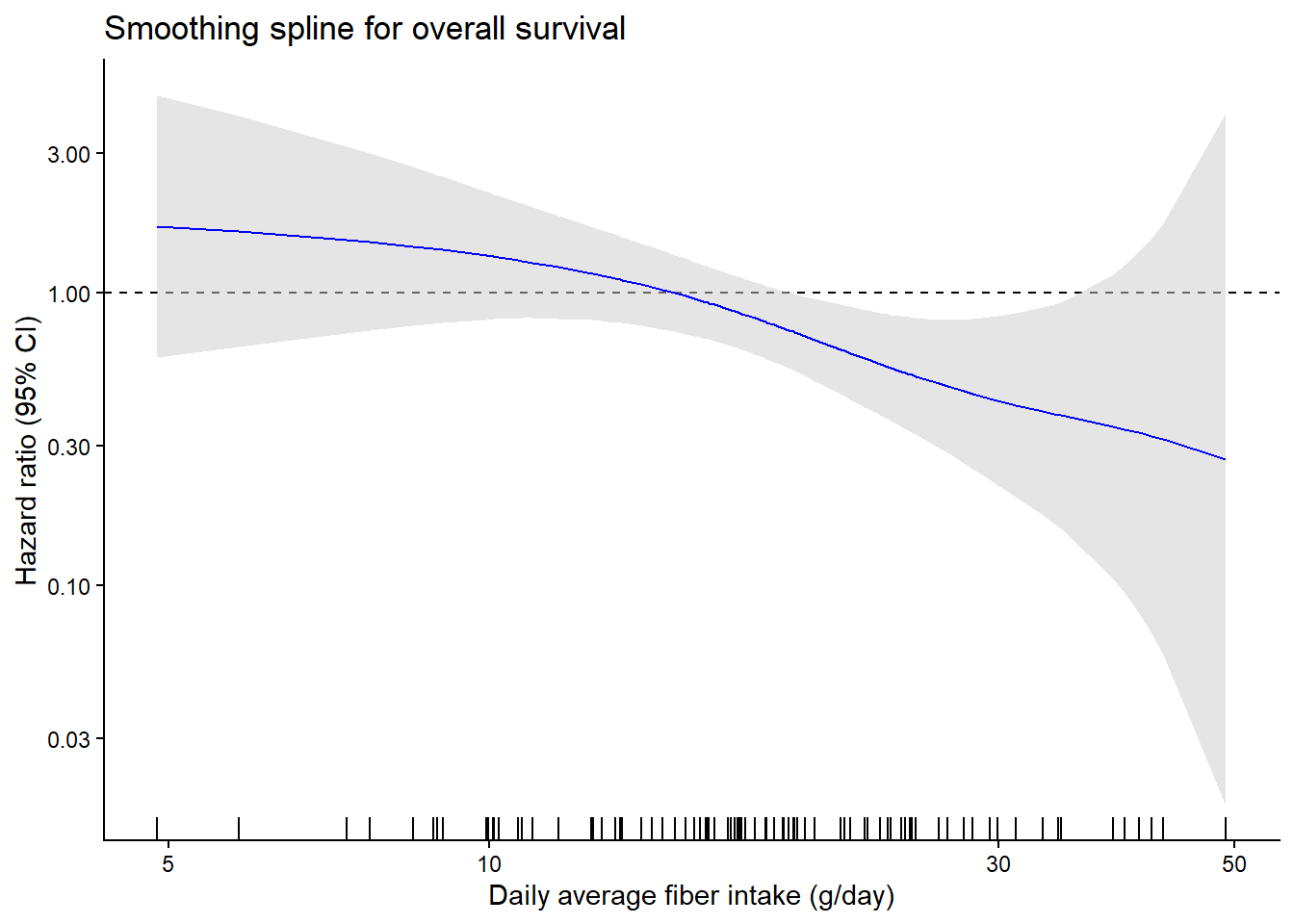
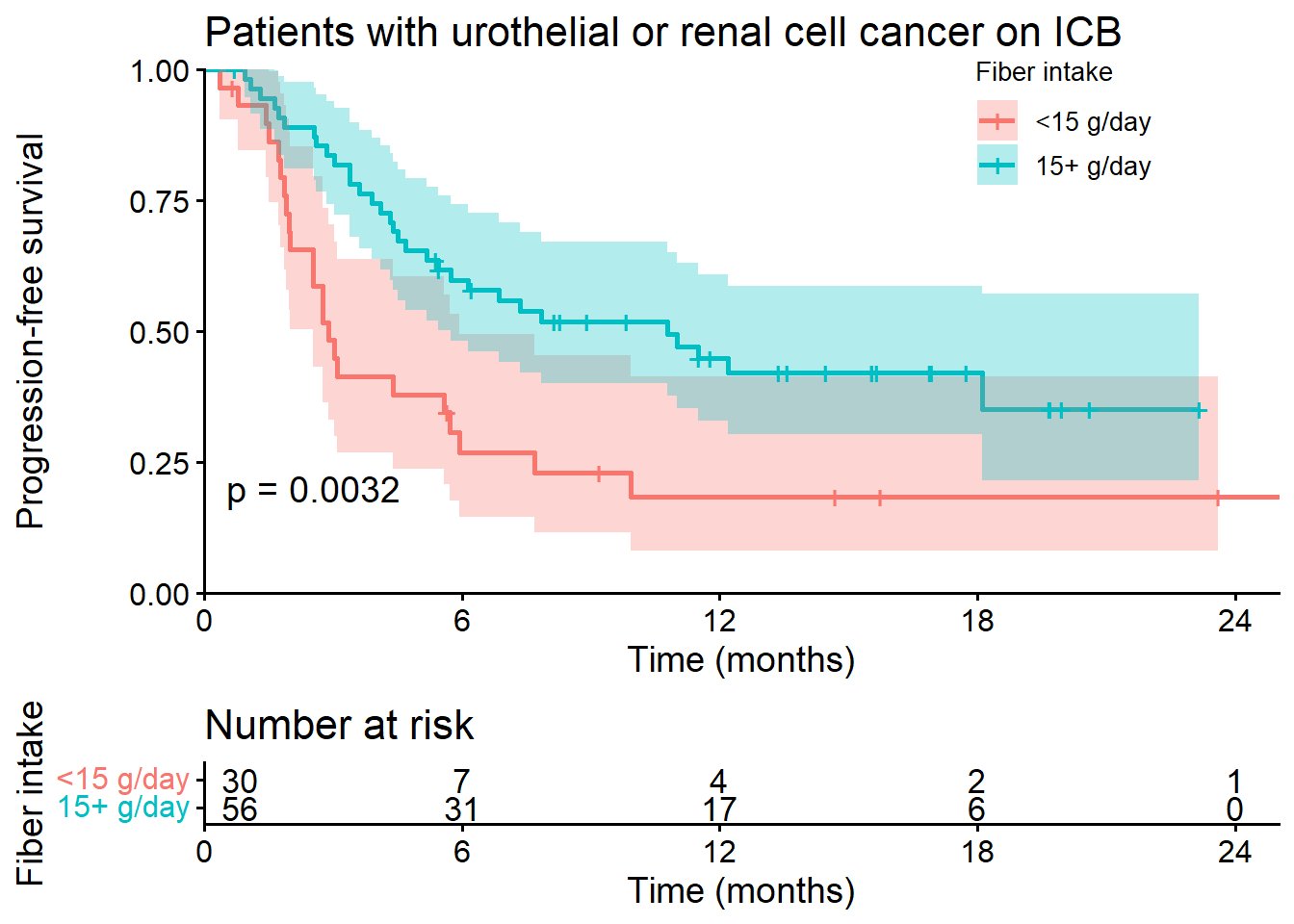
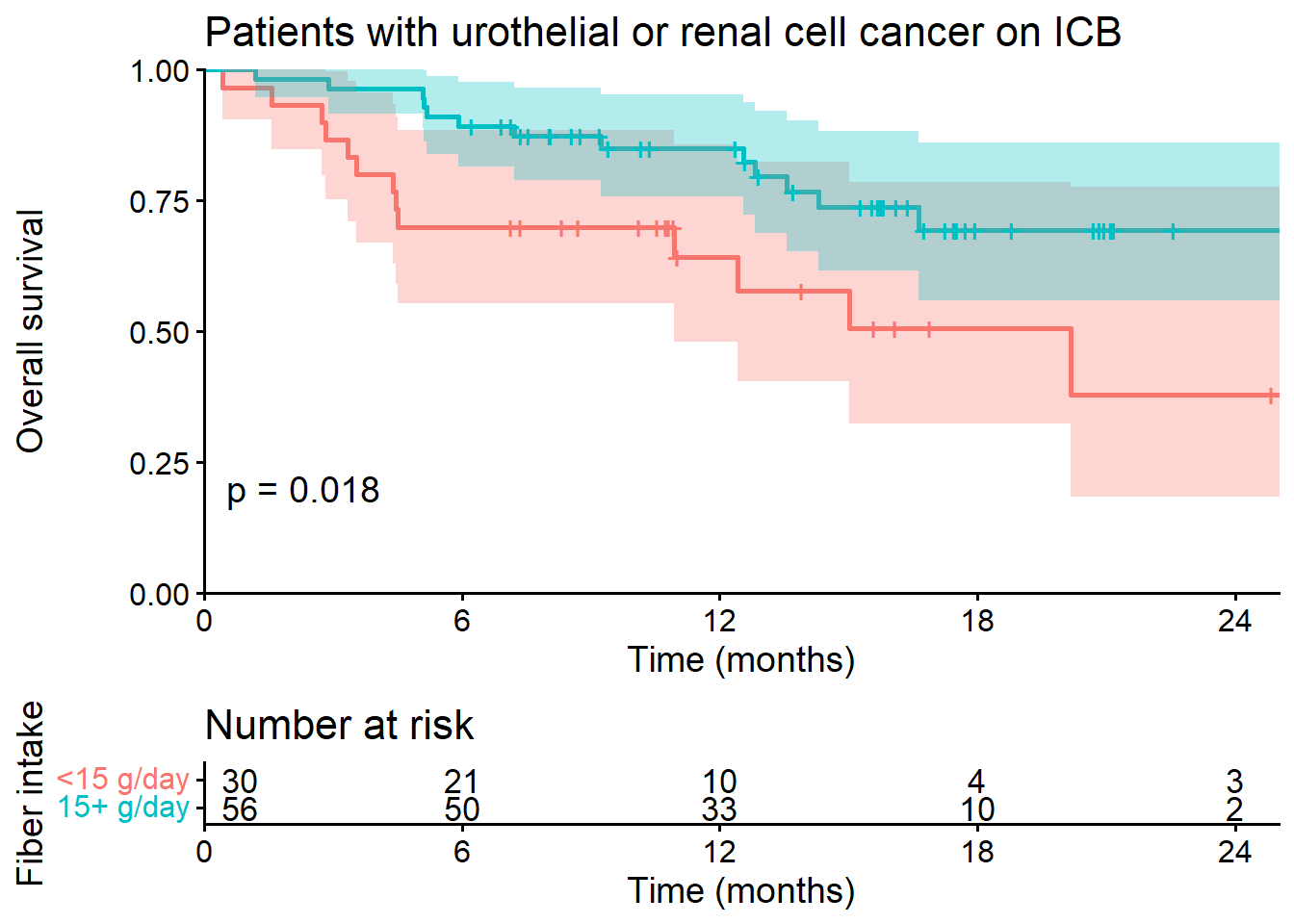
 Figure 1 - Hazard Ratio Analysis of Fiber Intake
Figure 1 - Hazard Ratio Analysis of Fiber Intake