[1] "blinded_subject_id" "id_int" "cohort"
[4] "age" "impact_tmb_score" "cpb_drug"
[7] "ecog" "best_overall_response" "tt_pfs_d"
[10] "pfs_event" "tt_os_d" "event_os"
[13] "TMPT" "WMS_SGPID" "SAMPID"
[16] "total" "samp_id" "identifier.y"
[19] "Shannon" "tx" "age10"
[22] "med_buty" "fouraminobutyrate" "gluatarate"
[25] "lysine" "pyruvate" Extended Figure 2 Gene Abundance and Hazard Ration, and Univatiate Analysis of Butyrate Pathway.
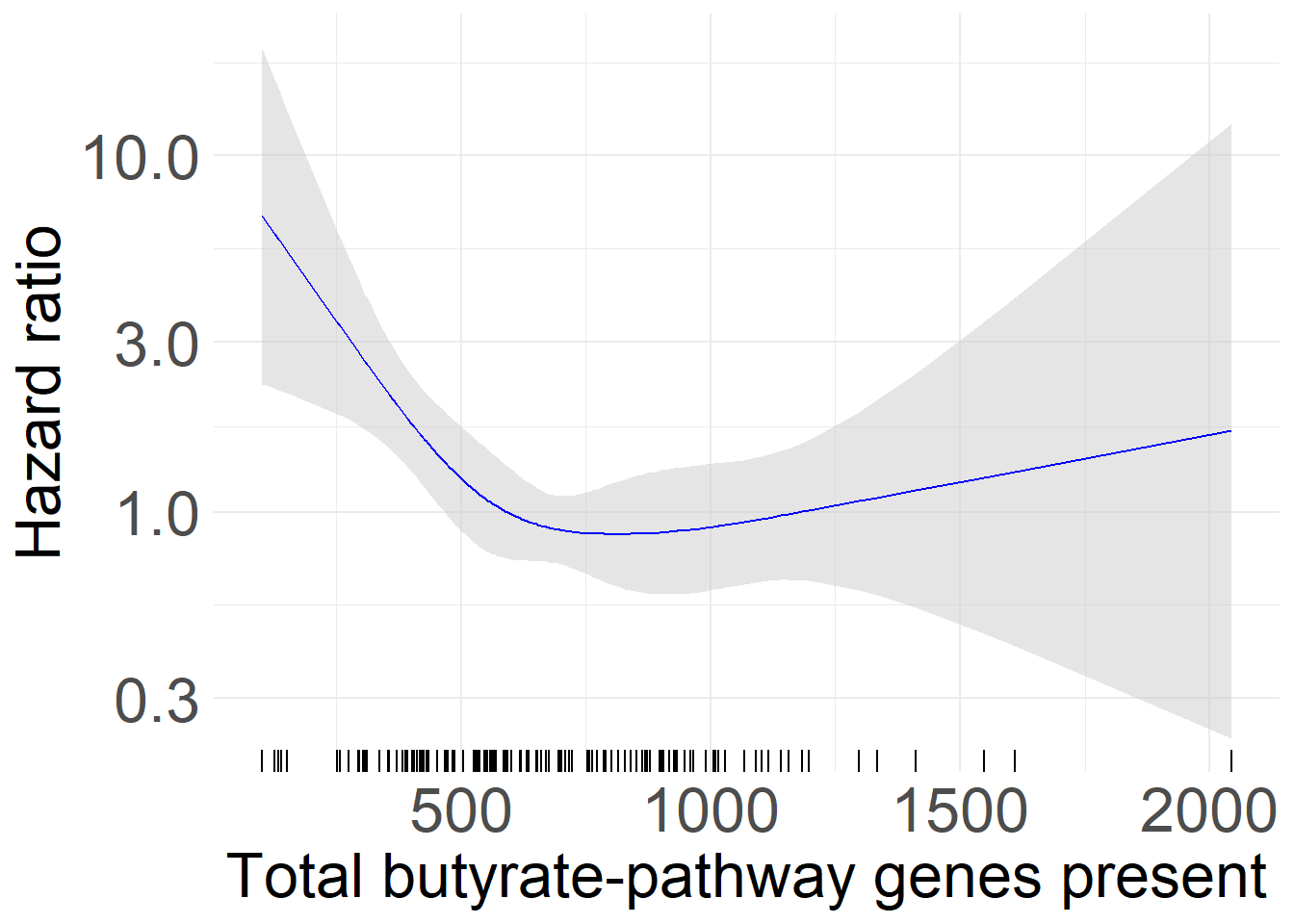
Panel A
Hazard Ratio of Overall Survival Evaluated with respect to Butyrate production gene RPKM.
os_butyrate <- coxph(Surv(tt_os_d, event_os)~rms::rcs(total,4)+age10+factor(cohort)+ecog,, data = acoa_df)
ptemp <- termplot(os_butyrate, se=T, plot=F)
buterm <- ptemp$total
center <- buterm$y[which(buterm$y == median(buterm$y))]
ytemp <- buterm$y + outer(buterm$se, c(0, -1.96, 1.96), '*')
exp_ytemp <- exp(ytemp - center)
spline_data <- data.frame(buty = buterm$x, Estimate = exp_ytemp[,1],
Lower = exp_ytemp[,2], Upper = exp_ytemp[,3])
ggplot(spline_data, aes(x = buty)) +
geom_ribbon(aes(ymin = Lower, ymax = Upper), fill = "grey80", alpha = 0.5) +
geom_line(aes(y = Estimate), color = "blue") +
geom_rug(sides = "b") + # Add rug plot at the bottom ('b') of the plot
scale_y_log10() + # Log scale for y-axis
labs(x = "Total butyrate-pathway genes present", y = "Hazard ratio") +
theme_minimal() +
theme(
axis.text = element_text(size = 24), # Adjust axis labels
axis.title = element_text(size = 24), # Adjust axis titles
plot.title = element_text(size = 24), # Adjust plot title
legend.text = element_text(size = 18), # Adjust legend text
legend.title = element_text(size = 20), # Adjust legend titleMetronidazole
strip.text.x = element_text(size = 20),
strip.text.y = element_text(size = 20)
) 
ggsave("svg/extended_figure_2_part_a_HR_tot_butyrate.svg", w = 6.5, h = 4) Panel B
Univariate model of OS as a function of total butyrate genes, and each major sub-pathway in the butyrate production gene pathways.
univ2 <- function(cox, k=1, l=1){
coef <- exp(coef(cox))
confint <- exp(confint(cox))
p <- summary(cox)$coefficients[,5]
out <- data.frame("HR"=round(coef,4), round(confint,4), "p"=round(p,9))
q <- nrow(out)
if(l== -1){out <- out[q,]
}else{out <- out[k:l,]}
return(out)
}
forest_os <- rbind(
univ2(coxph(Surv(tt_os_d, event_os)~log(total)+age10+cohort+ecog, acoa_df)),
univ2(coxph(Surv(tt_os_d, event_os)~log(pyruvate)+age10+cohort+ecog, acoa_df)),
univ2(coxph(Surv(tt_os_d, event_os)~log(lysine+0.0000001)+age10+cohort+ecog, acoa_df)),
univ2(coxph(Surv(tt_os_d, event_os)~log(fouraminobutyrate+0.000001)+age10+cohort+ecog, acoa_df)),
univ2(coxph(Surv(tt_os_d, event_os)~log(gluatarate+0.000001)+age10+cohort+ecog, acoa_df)))
forest_os %>%
rename_with(~ c("HR", "CI_low", "CI_high", "p"), .cols = everything()) %>%
rownames_to_column("scfa") %>%
ggplot() +
geom_vline(xintercept=1, lty=2, size=0.25)+
geom_point(aes(x=HR, y=fct_reorder(scfa, HR, .desc = T)))+
geom_linerange(aes(xmin=CI_low, xmax=CI_high, y=fct_reorder(scfa, HR)), size=0.5) +
scale_x_continuous(transform = "log", breaks=round(exp(seq(-2,1.5,0.5)),1)) +
theme_classic() +
coord_cartesian(xlim=exp(c(-2, 0.3))) +
labs(y = "") +
theme(
axis.text = element_text(size = 20), # Adjust axis labels
axis.title = element_text(size = 20), # Adjust axis titles
plot.title = element_text(size = 20), # Adjust plot title
legend.text = element_text(size = 18), # Adjust legend text
legend.title = element_text(size = 18), # Adjust legend titleMetronidazole
strip.text.x = element_text(size = 18),
strip.text.y = element_text(size = 18)
) +
theme(axis.text.x = element_text(angle = 90, vjust = 0.5, hjust=1))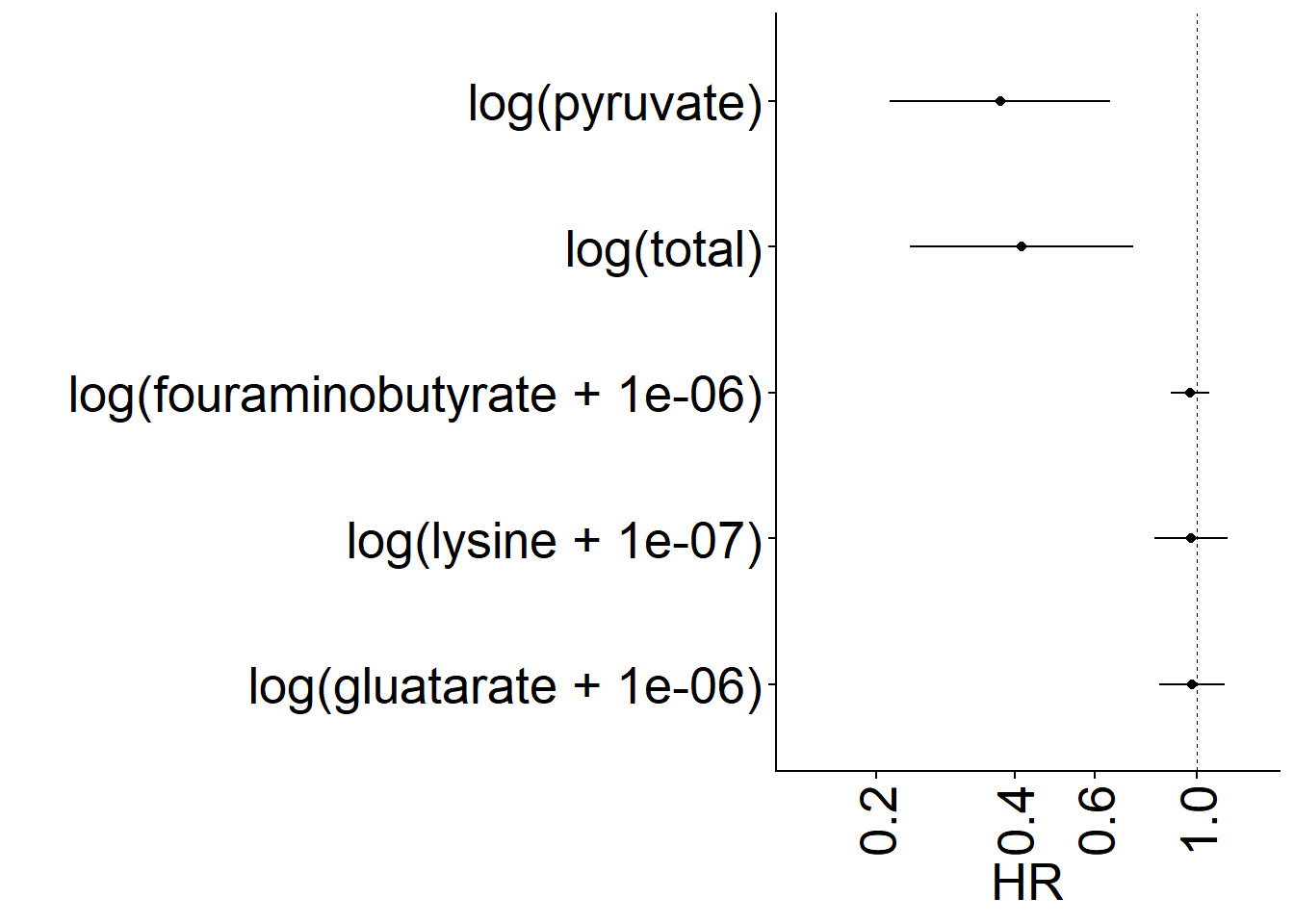
ggsave("svg/extended_figure_2_part_b_forest_plt.svg", w = 6.5, h = 4)