Relationship between relative abundance of top families and Acetyl-CoA pathway abundance
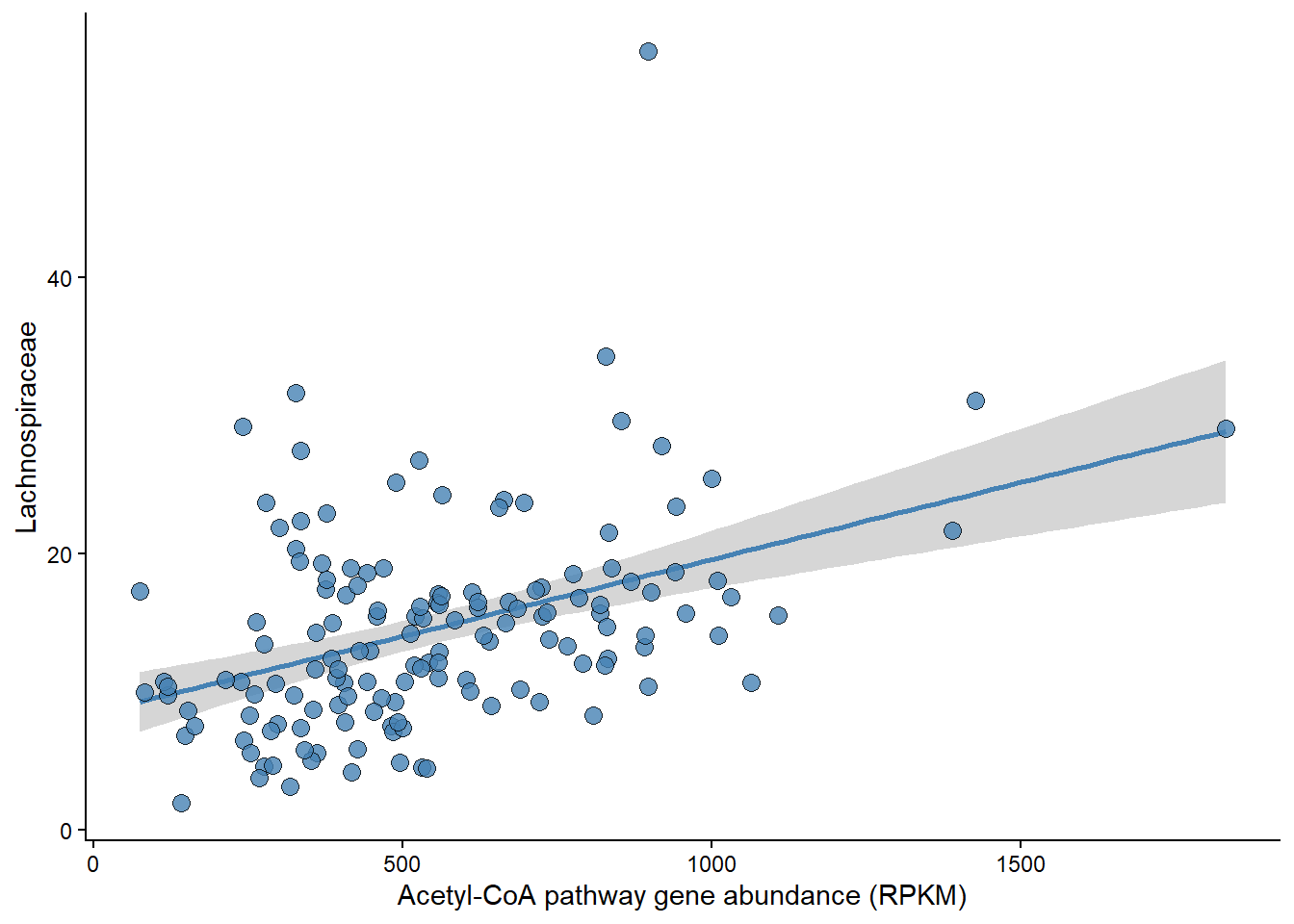
Panel A
Relationship between Rel Abundance of Lachnospiraceae and Acetyle-CoA pathway abundance.
plotting_ps %>%
filter(grepl("Lachnospiraceae", Family)) %>%
ggplot() +
geom_smooth(aes(x=acoa, y=Abundance), method=lm, color="steelblue") +
geom_point(aes(x=acoa, y=Abundance),
shape=21, size=3, fill="steelblue", alpha=0.8) +
theme_classic() +
labs(
x = "Acetyl-CoA pathway gene abundance (RPKM)",
y = "Lachnospiraceae"
)
ggsave("svg/extended_figure_4_part_a_Lachnospiraceae.svg", w=5, h = 5)
#|warning: false
#|label: p_value_panel_a
fit <- lm(Abundance ~ acoa, plotting_ps %>%
filter(grepl("Lachnospiraceae", Family)))
summary(fit)
Call:
lm(formula = Abundance ~ acoa, data = plotting_ps %>% filter(grepl("Lachnospiraceae",
Family)))
Residuals:
Min 1Q Median 3Q Max
-9.982 -4.407 -1.105 2.090 37.986
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 8.418826 1.232703 6.830 2.17e-10 ***
acoa 0.011170 0.001998 5.589 1.10e-07 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard error: 6.725 on 145 degrees of freedom
Multiple R-squared: 0.1773, Adjusted R-squared: 0.1716
F-statistic: 31.24 on 1 and 145 DF, p-value: 1.096e-07
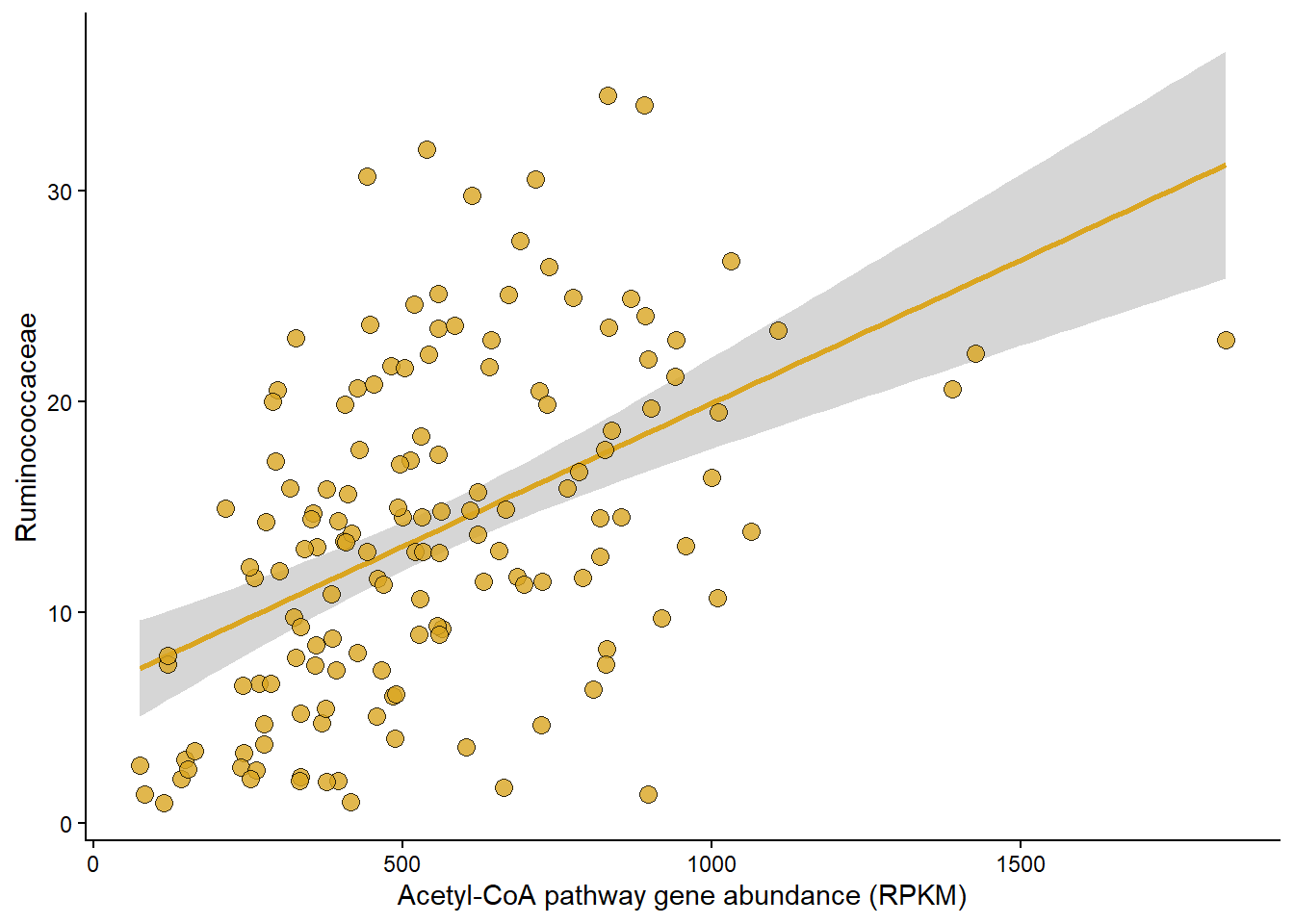
Panel B
Relationship between Rel Abundance of Ruminococcacae and Acetyle-CoA pathway abundance.
plotting_ps %>%
filter(grepl("Ruminococcaceae", Family)) %>%
ggplot() +
geom_smooth(aes(x=acoa, y=Abundance), method=lm, color="goldenrod") +
geom_point(aes(x=acoa, y=Abundance),
shape=21, size=3, fill="goldenrod", alpha=0.8) +
theme_classic() +
labs(
x = "Acetyl-CoA pathway gene abundance (RPKM)",
y = "Ruminococcaceae"
)
ggsave("svg/extended_figure_4_part_b_Ruminococcaceae.svg", w=5, h = 5)
#|warning: false
#|label: p_value_panel_b
fit <- lm(Abundance ~ acoa, plotting_ps %>%
filter(grepl("Ruminococcaceae", Family)))
summary(fit)
Call:
lm(formula = Abundance ~ acoa, data = plotting_ps %>% filter(grepl("Ruminococcaceae",
Family)))
Residuals:
Min 1Q Median 3Q Max
-17.1756 -5.2271 -0.5834 4.2678 18.3653
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 6.310352 1.285751 4.908 2.45e-06 ***
acoa 0.013612 0.002084 6.530 1.04e-09 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard error: 7.014 on 145 degrees of freedom
Multiple R-squared: 0.2272, Adjusted R-squared: 0.2219
F-statistic: 42.64 on 1 and 145 DF, p-value: 1.037e-09
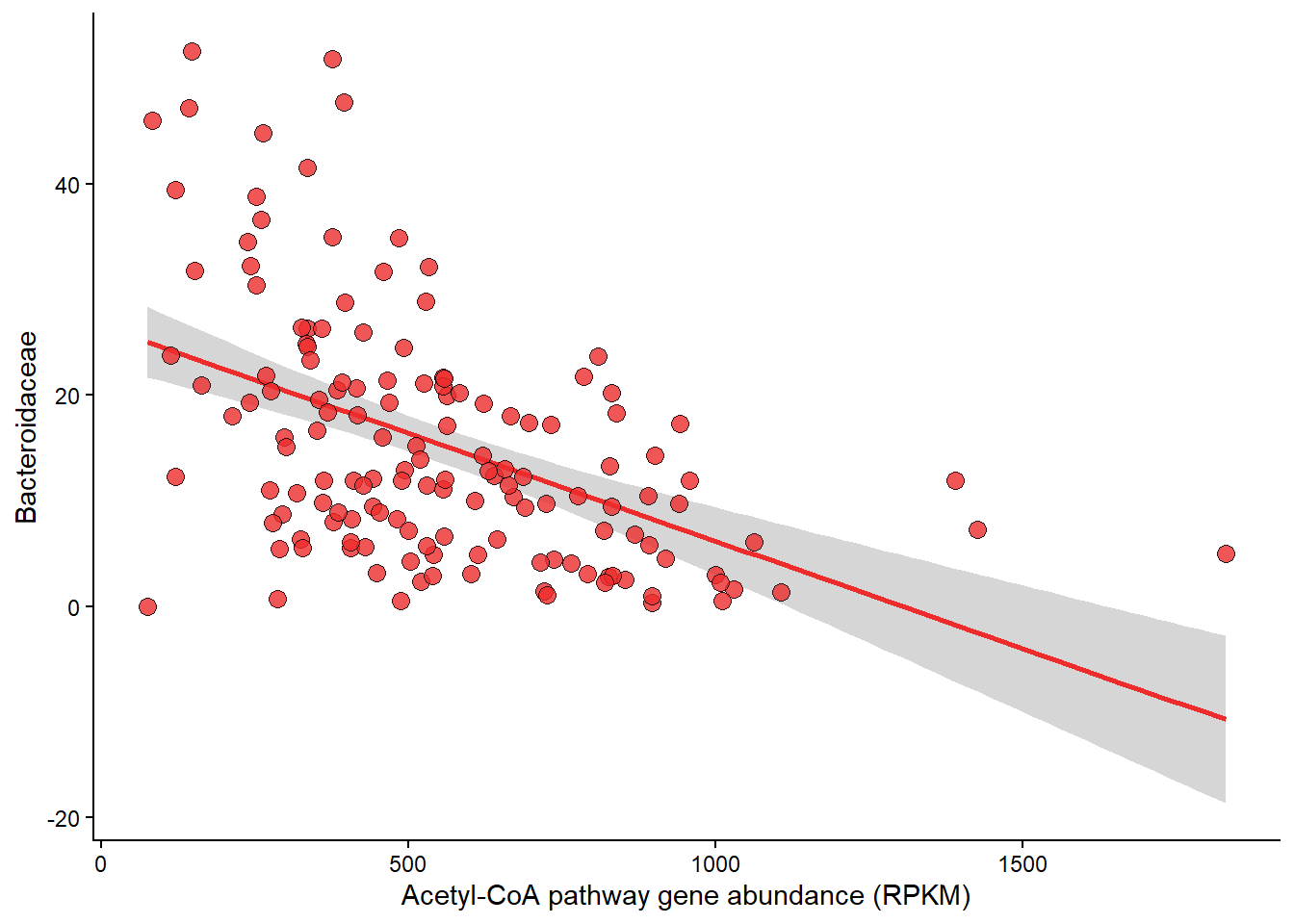
Panel C
Relationship between Rel Abundance of Ruminococcacae and Acetyle-CoA pathway abundance.
plotting_ps %>%
filter(grepl("Bacteroidaceae", Family)) %>%
ggplot() +
geom_smooth(aes(x=acoa, y=Abundance), method=lm, color="firebrick2") +
geom_point(aes(x=acoa, y=Abundance),
shape=21, size=3, fill="firebrick2", alpha=0.8) +
theme_classic() +
labs(
x = "Acetyl-CoA pathway gene abundance (RPKM)",
y = "Bacteroidaceae"
)
ggsave("svg/extended_figure_4_part_c_Bacteroidaceae.svg", w=5, h = 5)
#|warning: false
#|label: p_value_panel_c
fit <- lm(Abundance ~ acoa, plotting_ps %>%
filter(grepl("Bacteroidaceae", Family)))
summary(fit)
Call:
lm(formula = Abundance ~ acoa, data = plotting_ps %>% filter(grepl("Bacteroidaceae",
Family)))
Residuals:
Min 1Q Median 3Q Max
-25.059 -7.357 -1.563 5.631 32.901
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 26.589662 1.891717 14.06 < 2e-16 ***
acoa -0.020394 0.003067 -6.65 5.58e-10 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard error: 10.32 on 145 degrees of freedom
Multiple R-squared: 0.2337, Adjusted R-squared: 0.2284
F-statistic: 44.22 on 1 and 145 DF, p-value: 5.577e-10